About Me
Life is chemistry — but how does chemistry become life? This question sits at the heart of my research.
I am a protobiologist and biochemist working at the intersection of systems chemistry and molecular evolution,
combining experimental and computational approaches to understand how complex,
self-reproducing systems emerge from simple chemical building blocks.
I obtained my BSc in Biology from Ben-Gurion University, and my MSc in Molecular Biology from the Weizmann
Institute of Science. I conducted research in bacterial ecology, immunology, protein evolution and genomics before
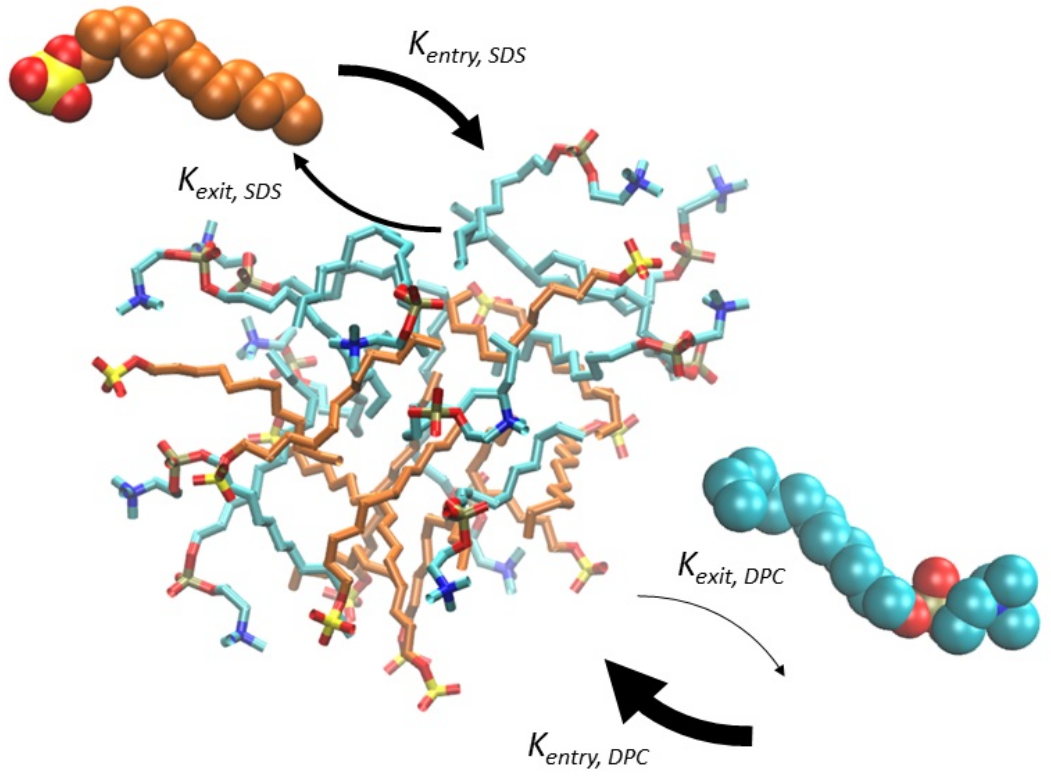
focusing on the origin of life — studying lipid-based self-reproducing catalytic networks in
the lab of Prof. Doron Lancet.
During my PhD in Chemistry at the University of Glasgow, under Prof. Lee Cronin, I further investigated
selection and evolution in chemical systems with Assembly Theory, developing analytical tools to uncover
causation at the molecular scale. I am now a postdoctoral researcher in the lab of Prof. Wilhelm Huck at Radboud
University, where I use AI-driven algorithms and automated platforms to navigate and control combinatorial chemical systems.
My broader scientific goals are to map the informational and functional landscape of living systems, uncover the
fundamentals of selection and evolution, and generate new living systems in the lab.
Outside of science, I spend my time cooking vegan food, sports climbing, hiking, and looking for the next adventure.

Research
-
2026 - present
Active Learning in Combinatorial Chemistry
Researcher in the lab of Prof. Wilhelm Huck.
Formulating AI-driven algorithms to empirically guide and control messy formose-based systems to uncover specific products and system dynamics, using automated experimental platforms and analytics. -
PhD in Chemistry | University of Glasgow
-
2022 - 2025
Selection in Complex Chemical Systems
Researcher in the lab of Prof. Lee Cronin.
Investigated Assembly Theory as a fundamental framework for the study of selection and evolution, through experimentation with advanced analytics and automated systems supported by theoretical work. -
MSc in Molecular Biology | Weizmann Institute of Science
-
2018-2022
Micellar Origin of Life
Researcher in the lab of Prof. Doron Lancet.
Researched the origin of life in a Lipid World scenario using advanced computational modelling and kinetic simulations of self-reproducing complex catalytic networks. -
2020-2020
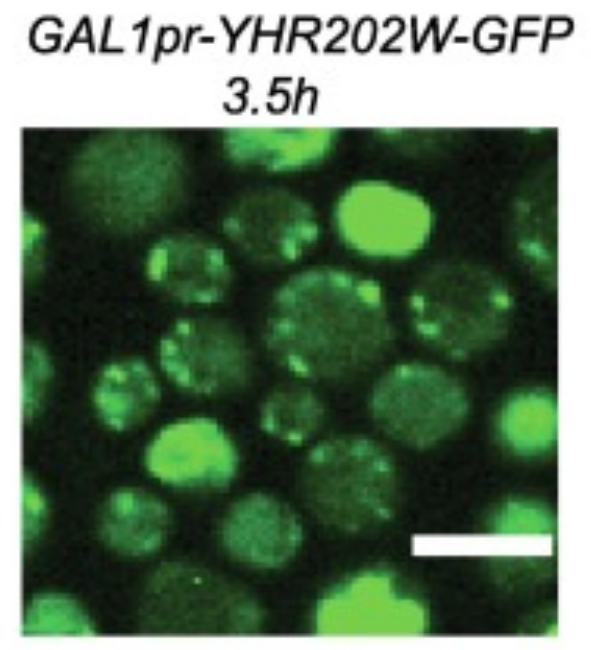
Functional Genomics
Research Intern in the lab of Prof. Maya Schuldiner.
Predicted the functionality of unexplored enzymes using advanced structural homology, and performed experimental validation. -
2019-2020
Protein Evolution
Research Intern in the lab of Prof. Dan Tawfik.
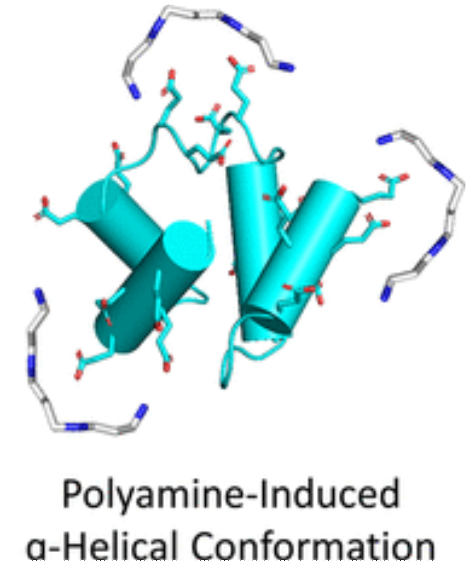
Examined the functional properties and folding capacities of a primordial RNA-binding protein candidate in the presence of polyamines. -
BSc in Biology | Ben-Gurion University
Ashalim Natural Sciences Honors Program
-
2017-2019
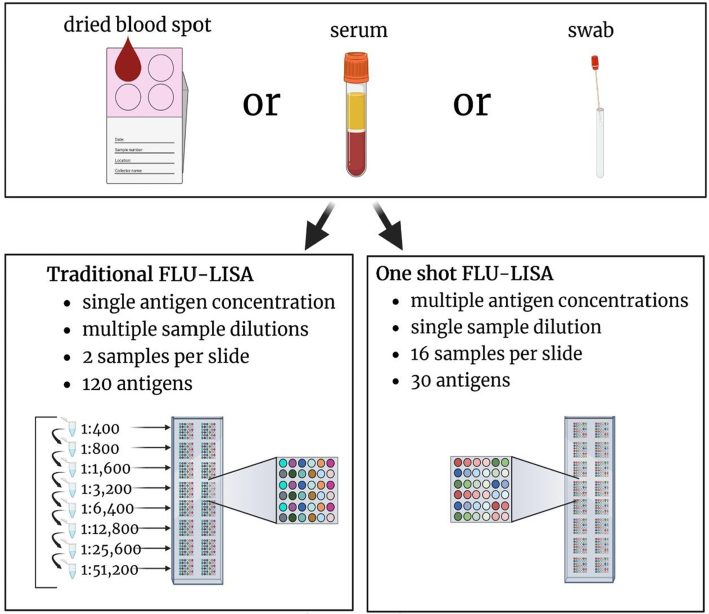
Avian Immunology
Research Assistant in the lab of Prof. Tomer Hertz.
Profiled the antibody repertoires of avian species and identified exposures to influenza and other viruses in relation to migration patterns of wild flocks. -
2018-2019
Microbial Ecology
Research Intern in the lab of Prof. Itzik Mizrahi.
Explored the dynamics of Horizontal Gene Transfer in microbial communities of ruminants.
Publications
Notable Projects:
2024
A Kahana, A MacLeod, H Mehr, A Sharma, E Carrick, M Jirasek, S Walker and L Cronin
arXiv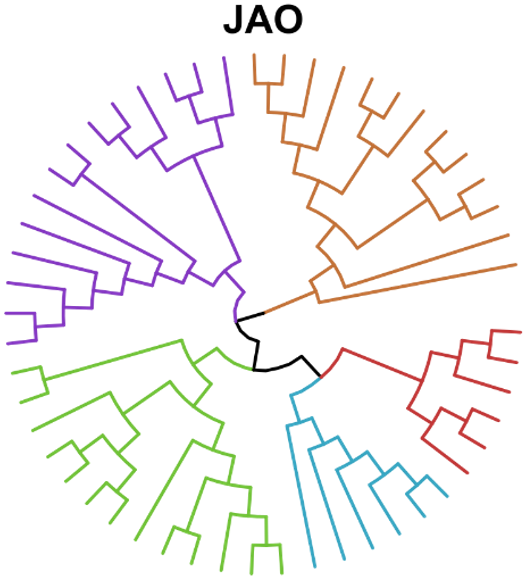
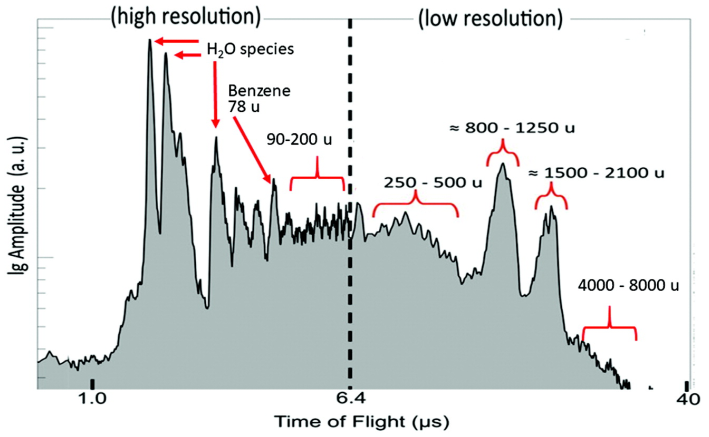
Can we reconstruct evolutionary relationships without knowing the identity of the molecules involved? We developed a method combining mass spectrometry and Assembly Theory that does exactly this — inferring how species are related by analyzing the complexity structure of their molecular mixtures, without needing to sequence DNA or identify individual compounds. Applied to a range of biological and non-biological samples, the approach successfully recapitulates the tree of life, tracks bacterial lineages, and opens the door to studying evolutionary relationships in chemical domains where sequence-based methods are notably limited.
2023
A Kahana, L Segev and D Lancet
CellRepPhysSci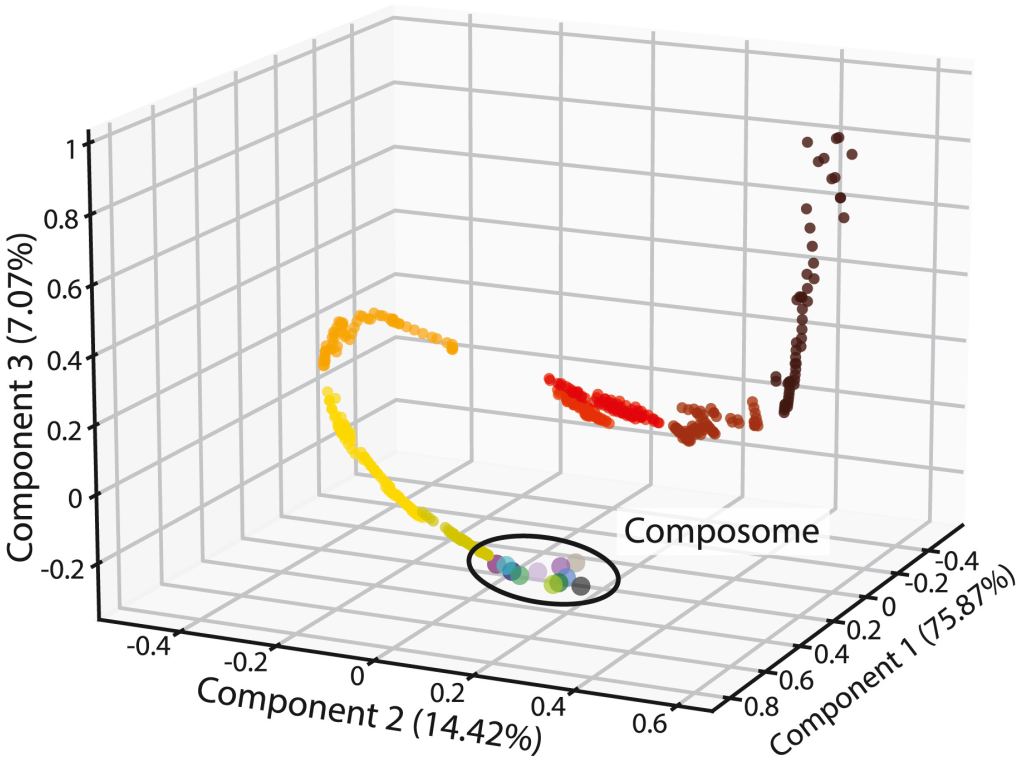
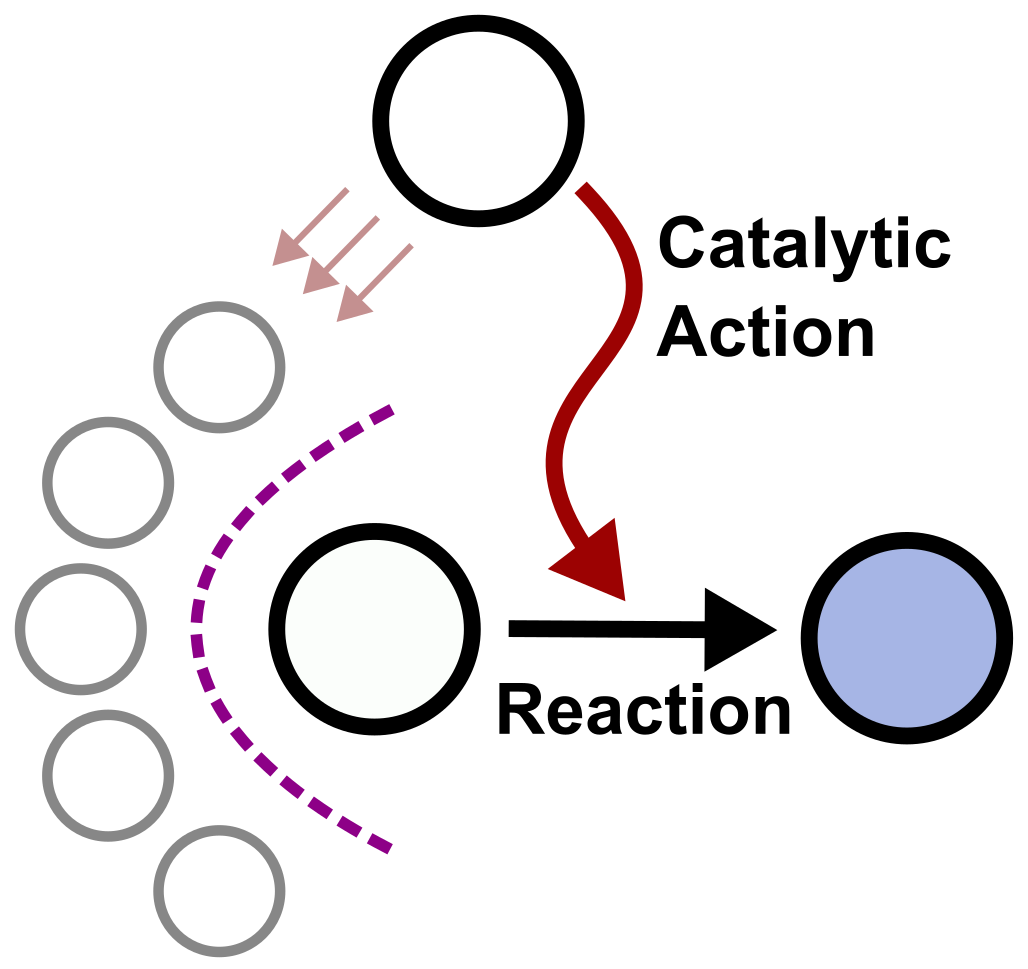
For life to emerge, chemistry had to become self-reproducing — but how likely is that to happen by chance? We explored this using the GARD model, a physicochemically rigorous simulation of self-replicating lipid networks. Self-reproducing compositions turn out to be rare in the space of possible chemistries, which would seem to make their spontaneous appearance unlikely. But we found a surprising saving grace: every self-reproducing state is a dynamic attractor of the catalytic network, meaning the system is naturally drawn toward these states over time. This dramatically increases the plausibility of life's emergence from disordered chemistry.
2021
A Kahana and D Lancet
NatRevChem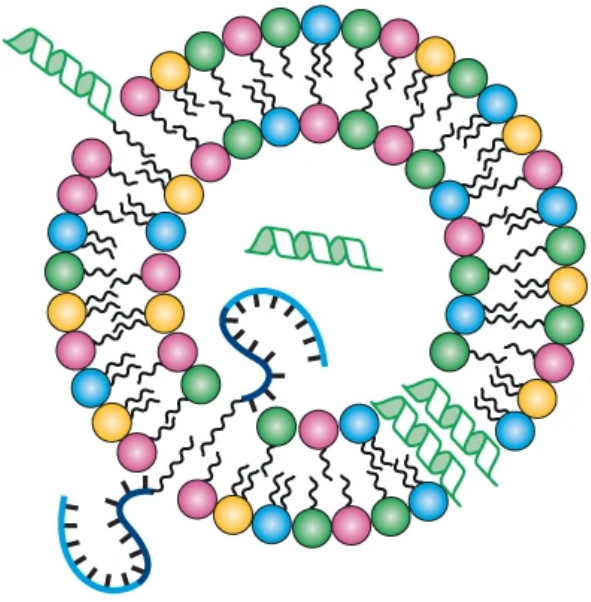
The conventional picture of the first protocells involves RNA or proteins enclosed in a lipid bilayer. But how did those biopolymers get there in the first place? We argue that self-reproducing lipid micelles came first — simpler structures capable of catalysis, growth, and compositional inheritance, requiring none of life's later molecular machinery. Drawing on experiments in micellar catalysis and our own computational modelling, we lay out a scenario in which lipid assemblies drove the earliest stages of molecular evolution, seeding the complexity that eventually gave rise to cellular life.
Other Selected Projects:
In Preparation
A Kahana, M Jirasek and L Cronin
2022
A Kahana, D Lancet and Z Palmai
Life2022
A Kahana*, N Cohen* and M Schuldiner
JMB2022
S Levy, M Abd Alhadi, A Azulay, A Kahana, N Bujanover, R Gazit, M A McGargill, L M Friedman and T Hertz
ICB2020
D Despotović, L M Longo, E Aharon, A Kahana, T Scherf, I Gruic-Sovulj and D S Tawfik
Biochemistry2019
A Kahana, P Schmitt-Kopplin and D Lancet
AstrobiologyFor a full list, see my Google Scholar profile.